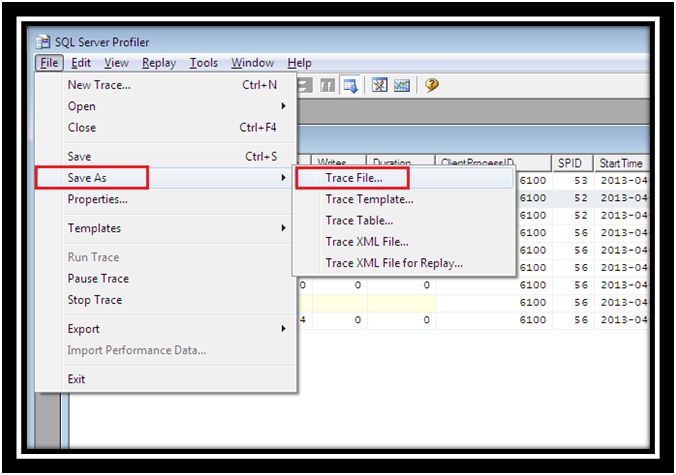
Patient characteristics according to HER2 mutation status. “4Peaks” was used to view and edit the sequence trace files ( ). The left half of Fig 3 shows the amplification curves of Eprobe-PCR, and the right half shows the electrogram of Sanger sequencing. The data were then transferred to Microsoft Excel (Microsoft, Redmond, WA, USA) and Cp values were evaluated.Įvaluation of the nine mutant tumors detected by Eprobe-PCR.Ĭomparison of three genotyping methods in all clinical samples.Ĭomparison of Eprobe-PCR and Sanger methods. (c) Cp (crossing point) values of two experiments (a) and (b) were calculated by the second derivative maximum method in the LightCycler480. Read reviews, compare customer ratings, see screenshots, and learn more about. The light blue line shows no amplification. The total copy number for each was adjusted to 10,000 copies per reaction. The blue line indicates MT only plasmid DNA at 10,000 copies per reaction, red: 1,000, green: 100, purple: 10, light blue: 1, orange: WT plasmid DNA, black: NTC (diluted water). (b) Sensitivity of 12-bp duplicated insertion detection in heterogenetic conditions. etl files so you must first open the file with Microsoft Message Analyzer and then export the results to a. Unfortunately WireShark cannot directly open. Traces appear sharper when viewed through 4Peaks compared to other chromatogram viewing. The blue line indicates MT only plasmid DNA at 10,000 copies per reaction, red: 1,000, green: 100, purple: 10, light blue: 1, orange: WT plasmid DNA, black: NTC. netsh trace stop Analyzing the capture: My personal preference is to use WireShark to process the results of netsh packet captures. 4Peaks boasts that it is able to render peaks better than anyone else. (a) Evaluation of mutated genome amplification. MT: HER2 12-bp duplicated insertion mutation type, WT: HER2 wild type, NTC: No template control (diluted water). Sensitivity of Eprobe-PCR for detecting HER2 12-bp duplicated insertion. Expect to get 500-700 bases of clean reliable DNA sequence. You should see individual, sharp and evenly spaced peaks like these. 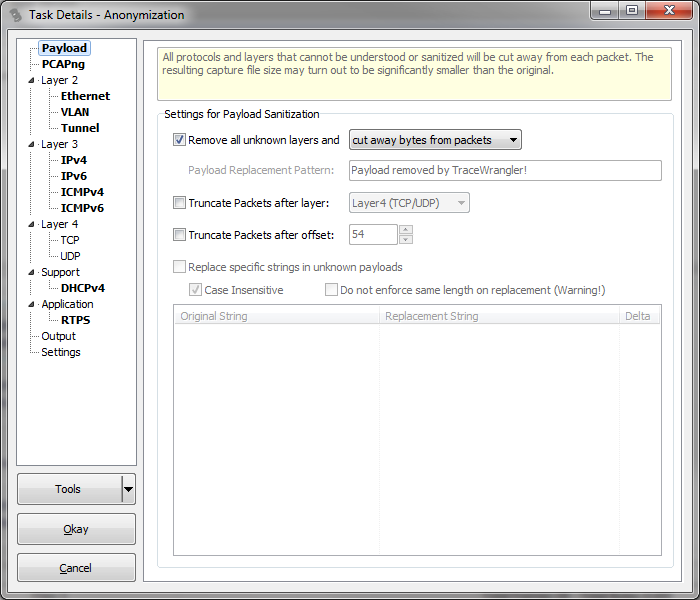
ab1 chromatogram file 4Peaks (Mac) SnapGene Viewer (Mac/PC) FinchTV (Mac/PC) Sequence Scanner (PC) Chromas (PC) 2. Read 1 user reviews and compare with similar apps on MacUpdate. You can use any of the following programs to view your. 4 peaks for Macintosh OSX is available at. SnapGene Viewer and NCBI STEP 1: Edit the Forward Trace File q Open SnapGene Viewer. For example, for User1 the TTD files would be stored here: Console. All of the trace output files are stored in the users document folder by default. The 3’ end-filled circle Eprobe shows the blocker that prevents primer extension during PCR. ab1 files can be viewed using the following programs for Mac and PC: Finch TV for PC and MAC OSX is. RUN files can be opened after they are recorded using File > Start debugging > Open trace file. The forward primer for detection of the mutant-type allele contains the full sequence of HER2 across the region known to be a frequent insertion site. 2013 (GRCh38/hg38) group: Genes and Gene Prediction track: All GENCODE. The orange box is the duplicated insertion. Schematic diagram of primers for the detection of the HER2 12-bp duplicated insertion by Eprobe-PCR. Please make your choice based on your computer platform and operating system.Primer sets and Eprobe design for HER2 mutation detection. You will find information about downloading, installing and using the software. 4Peaks is a DNA managing software enabling molecular biologists to view, edit and manipulate DNA sequence files. Click on the appropriate icon(s) to go to the respective Web page. Tools for Viewing Sanger Sequencing Data Sequence / Chromatogram Viewing SoftwareĪ number of free software programs are available for viewing trace or chromatogram files. Azenta Life Sciences Consumables & Instruments.Gene Synthesis & Cloning/Mutagenesis FAQs.CLIA Variant Confirmation (PCR + Sanger).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
